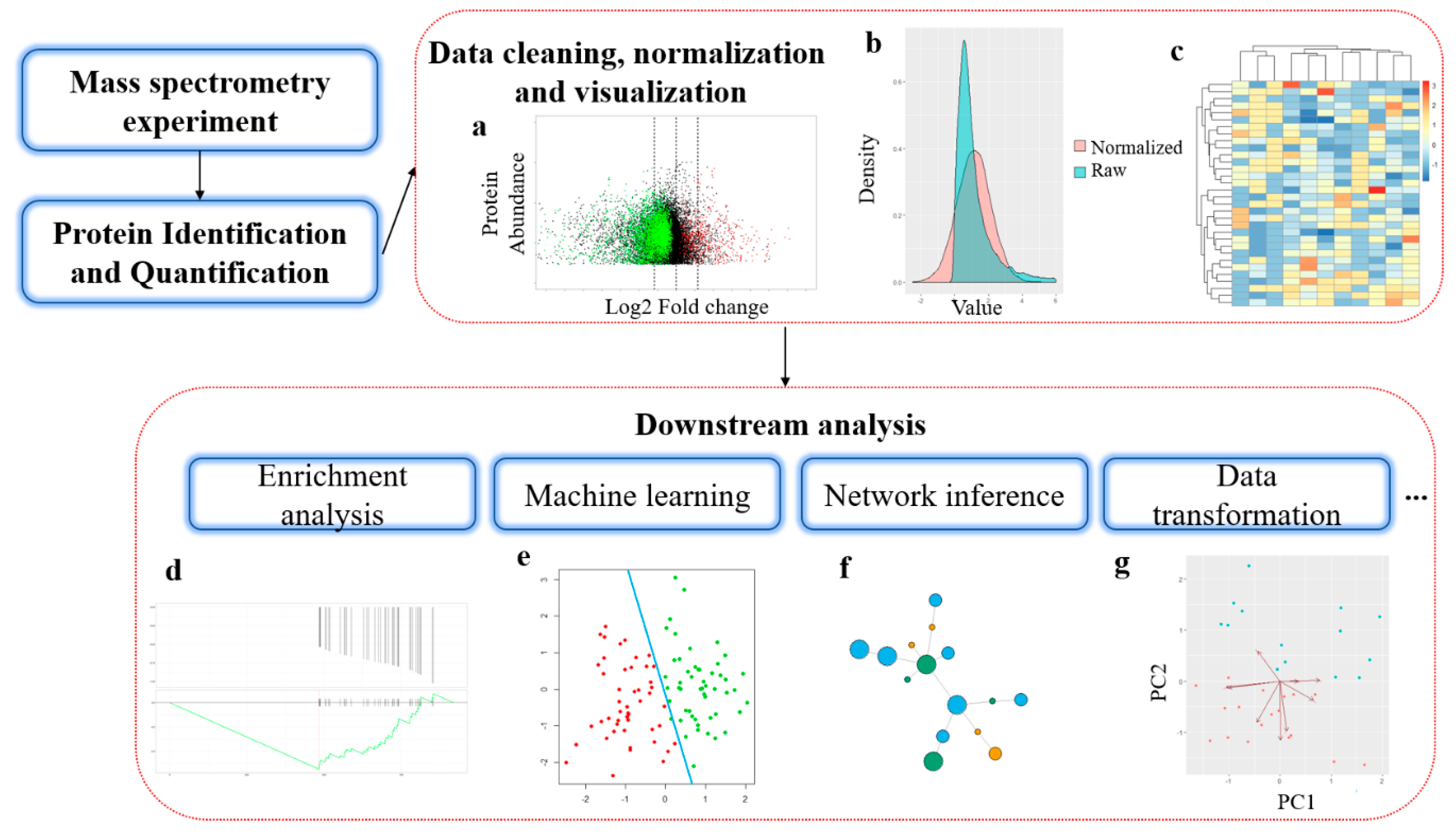
Simplifying MS1 and MS2 spectra to achieve lower mass error, more dynamic range, and higher peptide identification confidence on the Bruker timsTOF Pro | PLOS ONE

Tertiary Structural Motif Sequence Statistics Enable Facile Prediction and Design of Peptides that Bind Anti-apoptotic Bfl-1 and Mcl-1 - ScienceDirect

A calculated isotope distribution. The mass spectrum of a peptide or... | Download Scientific Diagram

The IDAES process modeling framework and model library—Flexibility for process simulation and optimization - Lee - 2021 - Journal of Advanced Manufacturing and Processing - Wiley Online Library

Ad hoc learning of peptide fragmentation from mass spectra enables an interpretable detection of phosphorylated and cross-linked peptides | Nature Machine Intelligence
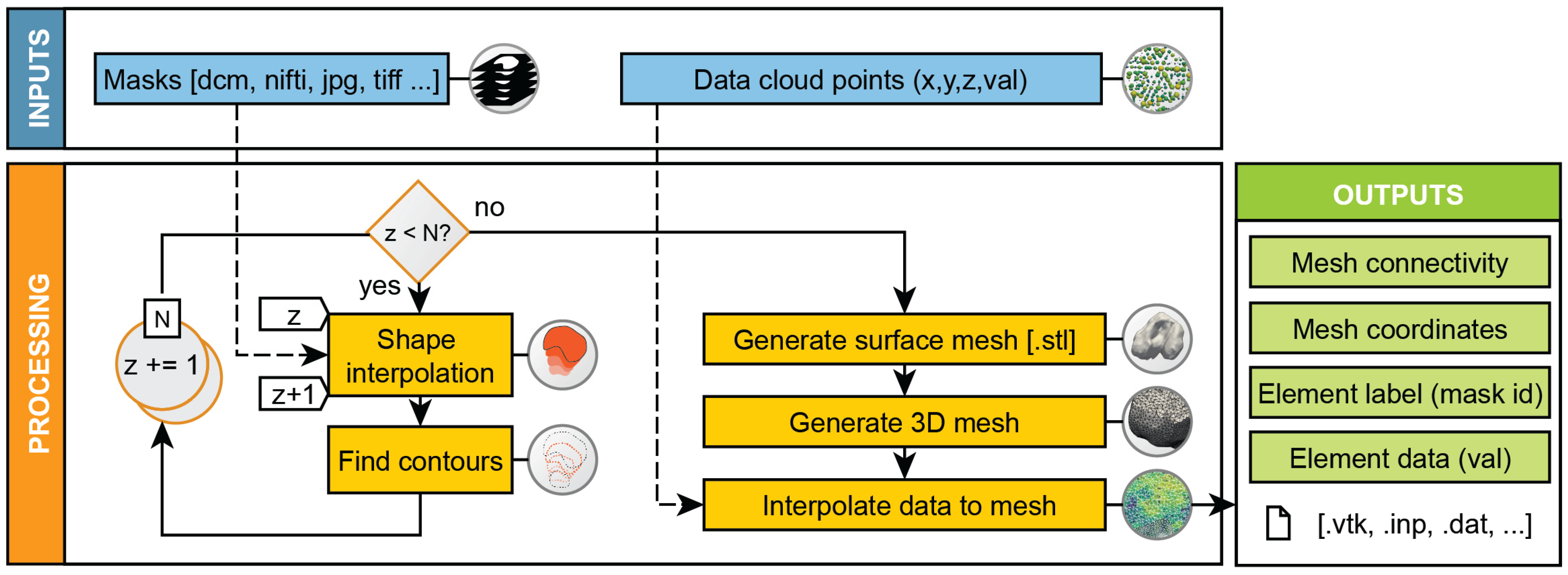
Applied Sciences | Free Full-Text | Im2mesh: A Python Library to Reconstruct 3D Meshes from Scattered Data and 2D Segmentations, Application to Patient-Specific Neuroblastoma Tumour Image Sequences

cgbind: A Python Module and Web App for Automated Metallocage Construction and Host–Guest Characterization | Journal of Chemical Information and Modeling
Simplifying MS1 and MS2 spectra to achieve lower mass error, more dynamic range, and higher peptide identification confidence on the Bruker timsTOF Pro | PLOS ONE

LC–MS peak assignment based on unanimous selection by six machine learning algorithms | Scientific Reports




![Using genetic programming to predict and optimize protein function [PeerJ] Using genetic programming to predict and optimize protein function [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2022/pchem-24/1/fig-2-full.png)